173
Continuum Mechanics
The macroscopic behavior and properties of engineering
materials depend on the microscopic characteristics
which are not accounted in typical continuum-based
models. In this project we operate on the atomistic scale,
and employ coarse-grained (CG) models as a computa-
tionally efficient means to study large numbers of atoms
over extended spatio-temporal scales.
A data-driven approach to CG has been proposed
within the the multidisciplinary joint project ‘Predictive
Materials Modeling’ with the Hans Fischer Senior Fellow
and TUM Ambassador, Prof. N. Zabaras (Viola D. Hank
Professor of Aerospace and Mechanical Engineering,
Director of the Computational Science and Engineering
Laboratory, University of Notre Dame, USA). The method
Coarse-Graining in Equilibrium Statistical Mechanics
developed makes use of generative probabilistic models
and employs a parametrized probabilistic mapping
from coarse-to-fine-scale descriptions which implicitly
defines the coarse variables. The coarse-variable model
is sequentially refined by adding features providing the
largest anticipated information gain. In the context of
peptide simulations, the dimension has been reduced by
a factor of 30, while still capturing the main properties and
revealing configurational similarities in the coarse space
as depicted in Figure 3a. Predictions that reconstruct the
full atomistic picture, are probabilistic and account for
uncertainties due to limited fine-scale simulation data and
information loss as a result of dimensionality reduction
(Figure 3b).
Well-established methods for the solution of
sochastic partial differential equations (SPDEs)
typically struggle in problems with high-dimen-
sional inputs/outputs. Such difficulties are only
amplified in large-scale applications where even
a few tens of full-order model runs are impracti-
cal. While dimensionality reduction can alleviate
some of these issues, it is not known which and
how many features of the (high-dimensional)
input are actually predictive of the (high-dimen-
sional) output. In this project, we advocate a
Bayesian formulation that is capable of perform-
Model-order and Dimension Reduction of Random Heterogeneous Media
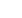
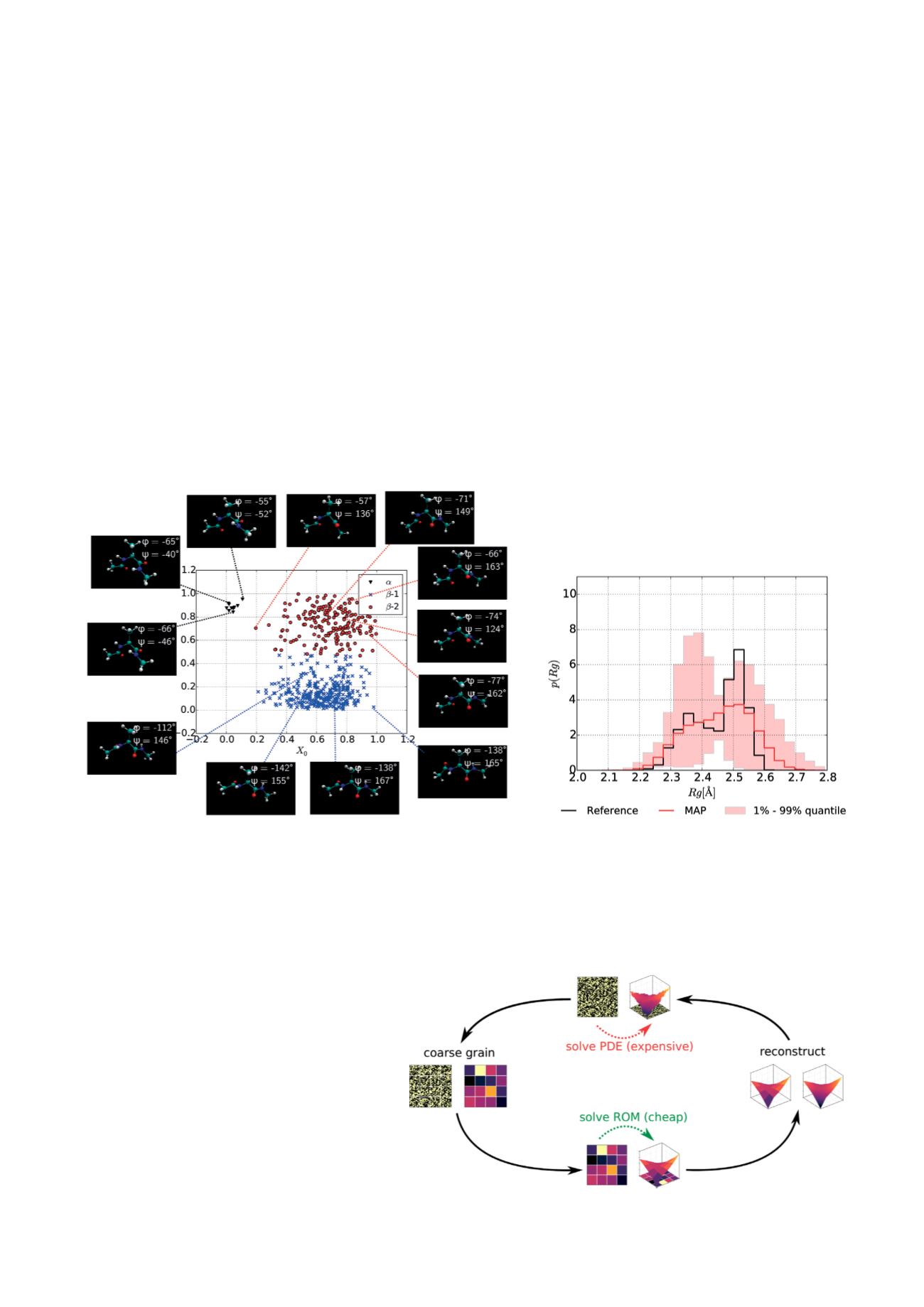
Figure 3: Predictive Coarse-Graining
(a) Representation of the training data in the reduced (two dimensional) latent space of
Alanine Dipeptide. Three clusters, corresponding to the characteristic configurational
modes (
a
,
b
– 1,
b
– 2), arise in the latent space.
(b) Probabilistic prediction of the deviation of the radius of
gyration from a common
a
– helix
Figure 4: Proposed framework



















